Hybrid Markov chain models for disease dynamics
From the early work of Daniel Bernoulli to the recent outbreak of Ebola, the mathematical modelling of infectious diseases is of fundamental importance to quantifying the threats that diseases pose to society. These models often strive to achieve either a high degree of accuracy or to be easily amenable to analysis by practitioners. Continuous-time Markov chain (CTMC) models are currently on the forefront of epidemic modelling because they capture a balance between these two criteria. However, modelling large populations with a CTMC model is usually computationally infeasible because it requires more computational power than is accessible to a majority of researchers.
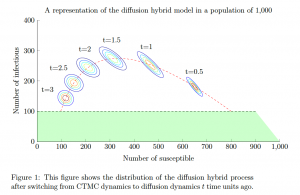
The aim of this project is to overcome this computational barrier by creating two novel hybrid models for approximating the CTMC model. To do so, we use the law of large numbers and central limit scalings of Thomas Kurtz and Andrew Barbour to create a so called fluid hybrid model and a diffusion hybrid model. Depending on its current state, the fluid hybrid process either has the dynamics of the CTMC or a particular ordinary differential equation. Similarly, the diffusion hybrid process either has the dynamics of the CTMC or a particular diffusion process. The figure below illustrates how the dynamics of the diffusion hybrid model are constructed when they are applied to the well-known susceptible-infectious-recovered (SIR) epidemic model. The hybrid models can be used to predict a number of important epidemiological quantities such as the distribution of the final epidemic size and the distribution of the duration of an epidemic. Or they can be used to improve the efficiency of computationally intensive algorithms such as Bayesian parameter inference.
Associated staff: Student — Nic Rebuli; Other — Prof. Nigel Bean